 
Jalview provides access to public services provided by the University of Dundee which are suitable for aligning up to 1000 sequences of at most 1000 residues each, and jobs are allowed to run for up to 1 h. Uniquely, coloring may be combined with conservation and consensus calculations to emphasize patterns of variation among conserved regions and to highlight different patterns of conservation among subgroups of sequences in the alignment.Ī range of alignment programs is offered via the Alignment submenu of the Alignment Window’s “Web Services” menu. A range of shading and coloring schemes allow common patterns to be highlighted according to standard properties for amino acids (such as hydrophobicity), or whether nucleotides are purine or pyrimidine derivatives.  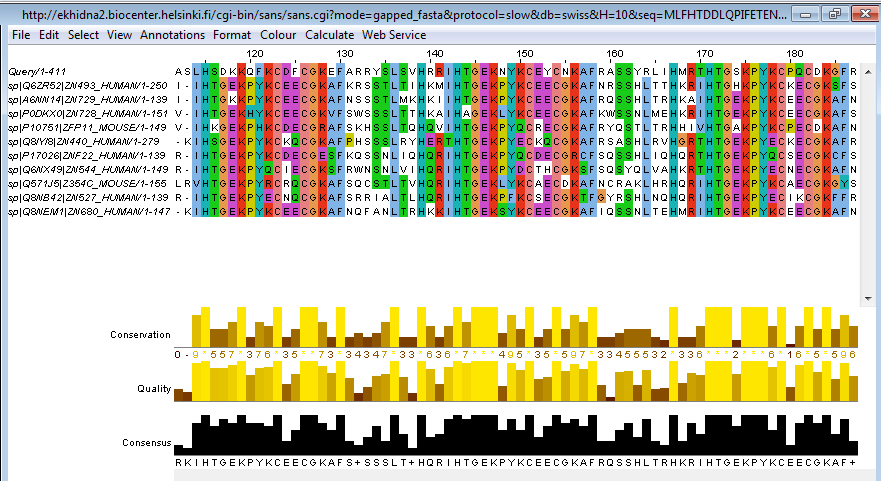
Aligned sequences may also be grouped either manually or by partitioning an associated tree which can be imported in a variety of formats or calculated using Jalview’s own tree algorithms. The alignment may be filtered to remove redundant sequences and to hide columns with low numbers of aligned residues. These create column Annotations that are shown as histograms under the alignment. Built-in analysis routines calculate the consensus for each alignment column and for proteins, the amino acid physicochemical properties. Access to a range of alignment programs is achieved directly within Jalview through web services, but Jalview also allows alignments generated by external programs to be imported. Sequences for alignment may be retrieved from public databases or loaded via a variety of common file formats. In this chapter, we describe the steps involved in a typical Jalview sequence alignment workflow, as depicted in Fig. The resulting alignments can be visualized and integrated with other information to further interpret them and create figures for publication. A core function is to make it easy to run state-of-the-art methods for multiple sequence alignment (MSA). 
The Jalview platform has many features for sequence analysis and visualization and is freely available both as a native “app” and single-page web application from its web site. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
